Paper
Combining rational design and computational tools in multi-parameter enzyme engineering to increase the fitness of a CYP152 peroxygenase for α-hydroxylation of fatty acids
S. Vrabl, K. Bangert, E. F. Semeraro, W. Kroutil
Green Chem 2026, 28, 7106–7116
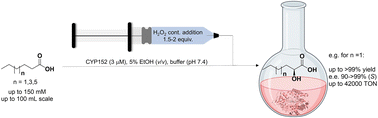
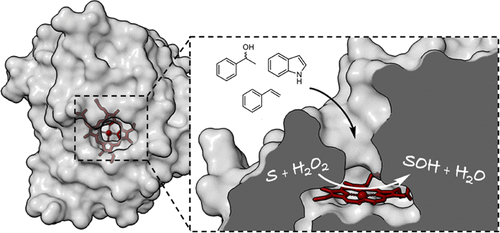
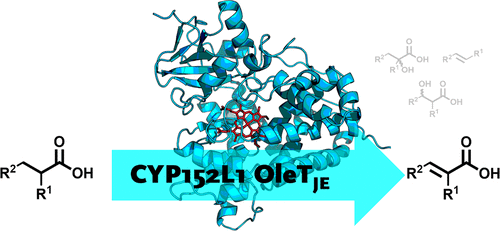
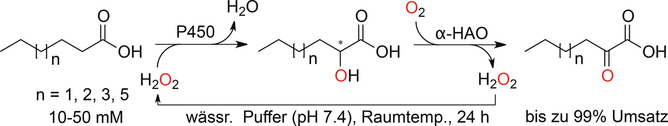
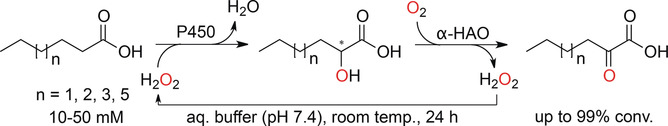
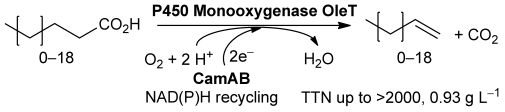
Enzymes of the CYP152 family have the potential to upgrade fatty acids to α-hydroxy acids by regio- and stereoselective hydroxylation using hydrogen peroxide as the oxidant and forming water as the sole by-product. To achieve such transformations with relevant productivity, a high tolerance towards hydrogen peroxide is required. By designing a minimal library targeting oxidation-prone residues with high solvent exposure and introducing mutations to improve expression/stability identified by a computational strategy (PROSS), the hydrogen peroxide tolerance of the CYP152 peroxygenase PO(SPα) was improved up to four-fold. Of the 12 generated enzyme variants, V3-P04 demonstrated the most pronounced improvements across the investigated parameters, exceeding the parent in terms of hydrogen peroxide tolerance, expression yields, and specific activity for the α-hydroxylation of octanoic acid. The superior performance of variant V3-P04 was further underlined in preparative-scale experiments for the functionalisation of heptanoic acid (50 mM), octanoic acid (150 mM), nonanoic acid (100 mM), and 9-decenoic acid (75 mM) where it reached turnover numbers unmatched by the wildtype enzyme of up to 48333.
doi: 10.1039/d6gc01139j



























 .
.